Our laboratory uses genetics and molecular biology to study pattern formation in the zebrafish embryo. The rapid development and simple anatomy of this teleost embryo, together with techniques for reverse genetics and a nearly complete genome sequence, make zebrafish a powerful molecular genetic system for studying the mechanisms of development. We are interested in how gene functions translate into cell behaviors and the formation of tissues and organs. We focus on three main areas:
1) Neural crest cell fate specification and formation of the craniofacial skeleton
2) Patterning of musculoskeletal attachments and roles of mechanical force in tendon development
3) Cell interactions and formation of the anterior-posterior axis of the nervous system
More recently we have begun to take an evolutionary approach using cichlid fish from Africa to map genes underlying natural craniofacial variation in the wild.
1) Craniofacial Development and Genetics
How do bones acquire their distinct sizes and shapes during development? Most bones of the skull are derived from cranial neural crest (NC) cells in the embryo, which migrate into the pharyngeal arches to form the facial skeleton. Defects in NC cells cause some of the most common birth defects in humans such as cleft palate, jaw and tooth defects and certain kinds of deafness. These cells have to find the appropriate segment (arch 1 versus 2) and acquire the proper dorsal-ventral (D-V) identity (lower versus upper jaw) within each arch. Over the years our work has focused on characterizing zebrafish mutants with craniofacial defects, which has led to the identification of novel molecular mechanisms underlying both the segmentation and D-V development of the facial skeleton. Our recent work has shown how Endothelin-1 (Edn1), Bone morphogenetic protein (Bmp) and Wnt signaling are integrated to determine craniofacial patterning (Alexander et al., 2011; Zuniga et al., 2011; Alexander et al., 2014).

Planar Cell Polarity and Skeletal Development
Stacks of chondrocytes are the basic building blocks of endochondral bones. We recently found that a mutant in REREa disrupts both the stacking and polarity of craniofacial cartilage. It appears to do so by interfering with planar cell polarity (PCP) signaling through the atypical cadherins Fat3 and Dachsous2 (Dchs2) (Le Pabic et al., 2014). This is the first evidence implicating Fat/Dchs signaling in skeletal cell polarity in vertebrates.

2) Cell Migration and Fate Specification
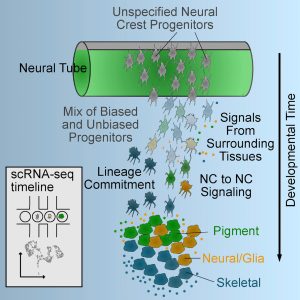
In addition to craniofacial cartilage and bone, NC cells also form peripheral neurons and glia, all of the body’s pigment cells, and many other derivatives. An over-arching goal of the lab is to understand how such migratory cells acquire their fates. Are they specified prior to migration or totally reliant on their migratory environments? Understanding this could help in the diagnosis and treatment of diseases that disrupt NC (e.g. cleft palate, neurofibromatosis, melanoma), including strategies for production of NC-like stem cells.
Neural Crest (NC) Cell Adhesion During Migration
How are migratory cell behaviors controlled by cell adhesion? NC cells rapidly alter distributions of adhesion molecules called cadherins at their surfaces during migration. Cadherins of the adherens junctions that hold epithelial cell together are downregulated in order for NC cells to undergo an epithelial-to-mesenchymal transition (EMT). Secreted Wnts induce some of these changes in NC adhesion and also promote specification of NC-derived pigment cells.
NC Cell Fate Specification
How do such highly migratory cells as NC cells acquire their diverse fates? Does this depend on their origins before migrating, migratory paths or the distinct environments in their ulimate destinations? To address this we recently performed single-cell RNA sequencing of subsets of migrating NC cells and showed that the earliest lineage decisions, at least in terms of transcriptional changes in gene expression, occur mid-migration. Not surprisingly, dynamic responses to Wnt signaling accompany these early fate decisions, and we use computational modeling to show how these correlate with early fate decisions (Tatarakis et al., 2021).
ECM Assembly and Muscle-Tendon Interactions at Myotendinous Junctions
Migrating cells also interact with ECM as they differentiate, a dramatic example of which occurs in myotendinous junctions (MTJs) that link muscles to bones. Cranial neural crest cells form the tendons of the head and these cells are thought to specify sites of muscle attachments to form a functional musculoskeletal system. We recently showed that a major component of the tendon ECM called Thrombospondin-4b (Tsp4b) maintains MTJ integrity in zebrafish and acts as a scaffold for other ECM components (Subramanian and Schilling, 2014). Overexpression of Tsp4, either fish or human, rescues muscle detachment and strengthens normal tendons, suggesting that it may be a potential therapeutic for tendon injuries. We are currently investigating the functions of the tendon determinant, Scleraxis, and several ECM proteins in the early development and maintenance of MTJs in zebrafish.

Anterior-Posterior Patterning and Retinoic Acid Signaling
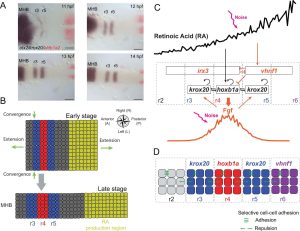
Another major area of research in the lab for many years focuses on the establishment of signaling gradients and their roles in cell fate decisions. The vitamin A derivative, retinoic acid (RA) is one such signal that is thought to act as a classic morphogen – specifying fates in a concentration-dependent manner – to pattern segments called rhombomeres along the anterior-posterior axis of the hindbrain. Our earlier work focused on the role of RA in determining segment identities and neuronal cell fates, while our more recent work focuses on RA degradation and the dynamics and robustness of the RA gradient itself.
Systems Biology: Spatial Dynamics in Retinoid Signaling
(White et al., 2007; White and Schilling, 2008; Cai et al., 2012; Zhang et al., 2012; Qiu et al., 2021)
Zebrafish as a Model for Studying Lens Physiology and Cataract Formation
Cataracts are the leading cause of blindness, due to the lens becoming opaque. One critical process for maintaining lens transparency is water permeability through channels consisting of Aquaporin-0 (Aqp0) proteins, but how Aqp0 is regulated and whether or not it has other functions remain unclear. In collaboration with Jim Hall’s lab in Physiology and Biophysics at UCI we recently found that zebrafish have two Aqp0s, only one of which acts as a water channel in permeability assays (Froger et al., 2010; Clemens et al., 2013). Interestingly, loss of function of either one causes cataracts in larval fish, and we are currently investigating other possible functions for Aqp0 (e.g. adhesion) that might be necessary for lens cell transparency.